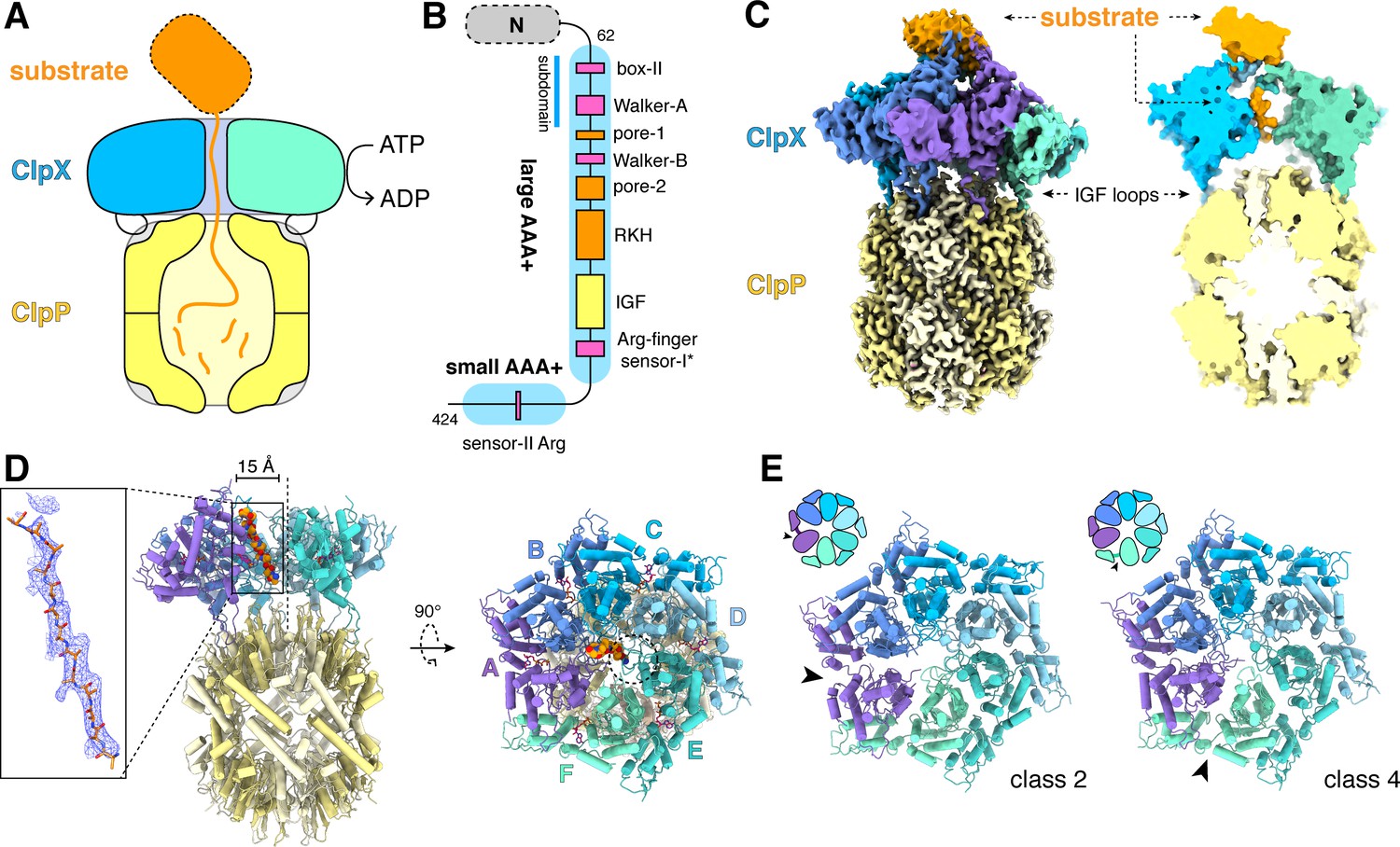
RCSB PDB - 3FES: Crystal Structure of the ATP-dependent Clp Protease ClpC from Clostridium difficile

Plantae | The CLP and PREP protease systems coordinate maturation and degradation of the chloroplast proteome in Arabidopsis thaliana (New Phytol.) | Plantae

ATP-Dependent Proteases Degrade Their Substrates by Processively Unraveling Them from the Degradation Signal: Molecular Cell